Magnetic micro-particle immune separation combined with enzyme linked immunosorbent assay for quick detection of Salmonella typhimurium in raw eggs
Cole Guyer, Shiquan Tao and Maya Alex
1Maya Alex, School of Medicine, University of Texas at El Paso, El Paso, TX 79968, USA
2Department of Mathematics, Chemistry and Physics, West Texas A&M University, Canyon, TX 79016, USA
- *Corresponding Author:
- Shiquan Tao
Department of Mathematics, Chemistry and Physics, West Texas A&M University, Canyon, TX 79016, USA
Tel: 8066512539
E-mail: stao@wtamu.edu
Received date: December 23, 2015; Accepted date: January 05, 2016; Published date: January 10, 2016
Citation: Tao S, Guyer C. Magnetic Microparticle of Combined with Enzyme Linked Immunosorbent Assay for Quick Detection of Salmonella typhimurium in Raw Eggs. Nano Res Appl. 2016, 2:1.
Abstract
An immunomagnetic separation (IMS) method was developed for separating Salmonella typhimurium bacteria from large-volume samples of raw eggs. An egg was homogenized with a blender. The homogenized egg material was diluted with DI water to make a 200 mL sample mixture. Anti-Salmonella typhimurium antibody coated magnetic micro particles (MMP) were used to trap the Salmonella bacteria in the sample mixture. The Salmonella-trapped MMP were separated from the egg sample matrices by using a large magnet. An enzyme-linked immunosorbent assay method was adopted and revised for detecting the bacteria trapped onto the MMP. A horseradish peroxidase (HRP) labeled anti-Salmonella typhimurium antibody was used to label the trapped bacteria, and a SureblueTM solution was used as a substrate. The color compound resulted from HRP-catalyzed reaction was detected with UV/Vis absorption spectrometry using a 1 cm sample cell. This simple method can detect 1.4*107 Salmonella typhimurium cells in one raw egg (7.0*104 Salmonella typhimurium cells/mL in a sample mixture) without any preenrichment. The results presented in this paper demonstrate the feasibility of using IMS for separating bacteria from large volume complex samples, which could be adopted for detecting bacteria in other type samples in food safety inspection.
Keywords
Salmonella typhimurium; Immunosorbent assay; Raw eggs; Food safety
Introduction
The quick detection of microbial pathogens in food materials has continuously been a challenge to the scientific community as well as to agricultural/food industries [1,2]. This challenge arises from following two facts: 1) the method has to be very sensitive, capable of detecting <10 pathogenic bacteria (e.g., E. coli O157:H7, Salmonella typhimurium), because these bacteria can cause severe disease, even death, even if only less than 10 bacteria are ingested. 2) food materials/products have very complex sample matrix composition. The sample matrix interferences prohibit the direct application of the most sensitive detection technologies, such as laser induced fluorescence and mass spectrometry which have the capability of detecting even a single molecule [3,4], for food sample analysis [5,6].
Traditional methods for food safety inspection employ microbiological approaches for detecting such pathogenic bacteria in food materials/products. These microbiological methods are very sensitive, can detect pathogenic bacteria down to single digit colony-form-unit per milliliter (CFU/mL) sample solution. Therefore, USDA Food Safety Inspection Services and US Food and Drug Administration (US FDA) recommend these methods as standard methods for food safety inspection [7,8]. Test results obtained with these microbiological methods are trustable, usually with no false response. However, these methods involve long-time enrichment processes (pre-enrichment, selective enrichment, plating on differential agar media) in order to increase the number of bacteria for detection. It usually takes 4-7 days to obtain a presumptive result [7,8]. The long waiting-time feature adds a high cost to agricultural products related to storage and freezing [9]. In addition, food quality also deteriorates during the long storage period. Therefore, there is a pressing need of quick detection technologies for food safety program.
Immunomagnetic separation (IMS) is a new development in bioanlytical chemistry for separating target biospecies from a sample solution. IMS uses antibody coated micro magnet particles (MMP) to trap target bacteria. Due to their magnetic property, the bacterium-trapped MMP can be easily separated from a complex sample solution by using a magnet [10-15]. In a recent development, IMS has also been integrated with a microchip technique for trapping Salmonella typhimurium from food samples on microchannels [16]. The trapped bacteria were labeled with a fluorescence reagent conjugated antibody. This provides a possibility of using a fluorescence microscope, which is broadly available in biotechnology laboratories, to count the limited number of bacteria. IMS is quick, convenient, selective and highly efficient for separating target bacteria from samples of complex matrices. In addition, this separation process does not involve any sophisticated equipment, which makes it useful for field applications. In addition to separation, IMS can also be considered as a concentration process [17,18]. Target bacteria can be concentrated from a large quantity (several hundred mL) sample to sub-milligrams MMP, and then be dissolved into submL solution. A concentration factor in the range of hundreds can be achieved.
In most reported IMS for food analysis works a pre-enrichment process is employed [13,15,16,19-21]. Food material/product samples were added to broth solutions in order to increase the number of bacteria for detection. The IMS separation method was then used to separate bacteria from pre-enrichment samples. In such cases, the sample matrixes in the time IMS was carried out are the broth solutions, which are relatively simple. The original sample matrix, such as egg white/yolk, meat, proteins in milk samples,does not have much effect on IMS separation process.
In food inspection programs, direct analyzing food samples without pre-enrichment is very attractive. Therefore, the focus of this work is investigating the feasibility of using IMS to directly separate bacteria from raw egg samples. Salmonella typhimurium was used as a target analyte because this bacterium is the most frequently reported problem with egg products. An enzymelabeled immunosorbent assay (ELISA) method was adopted and revised for detecting the Salmonella cells trapped onto the MMP. The whole analysis process takes less than 5 hours. With UV/Vis optical absorption spectrometry as a detection method, the developed IMS-ELISA method can detect 1.4*107 Salmonella typhimurium cells in one raw egg. It is believed that with the adaptation of more sensitive detection method, such as laser induced fluorescence [3,4], chemiluminescence [20], liquid core waveguide optical spectrometry [21,22], which is the focus of our further work, it is possible to achieve detecting of bacteria in single digit number in a raw egg sample.
Experimental
Materials and chemicals
A phosphate-buffered saline (PBS, 10X Phosphate-buffered Saline) solution from KPL Inc. (Gaithersburg, MD, USA) was 10- time diluted according to the maker’s instruction. The diluted PBS buffer solution was used in diluting/preparing reagent solutions and washing MMP in ELISA process. Anti-Salmonella typhimurium antibody coated MMP (BacTrace® Anti-Salmonella CSA-1 magnetic beads, MMP concentration>109 beads/mL, bead size is approximately 2 μm) from KPL Inc. were used for trapping Salmonella typhimurium bacteria from raw egg samples. A heatkilled Salmonella typhimurium cell sample (BacTrace® Salmonella typhimurium Positive Control, approximately 5 × 109 cells/mL in 1 mL of 2% w/v wet packed cells in dextran solution, KPL Inc.) was used in this work as a stock standard sample. A standard sample of 2.5*108 Salmonella typhimurium cells/mL was prepared by 20 times diluting the stock standard bacterial solution with the diluted PBS buffer solution. A horseradish peroxidase (HRP) labeled anti- Salmonella typhimurium antibody (BacTrace® Anti-Salmonella, CSA-1 Antibody, Peroxidase-labeled) was purchased from KPL, Inc. The enzyme labeled antibody material was dissolved in a 50% glycol solution as recommended by the maker to make a 0.1 mg antibody/mL solution. This antibody solution was further diluted with PBS buffer to make a 1 μg antibody/mL solution, which was used in the ELISA process of this work. SureBlueTM TMB Microwell peroxidase substrate from KPL Inc. was used as received as an enzyme substrate. TMB BlueSTOPTM solution from KPL Inc. was used as received to stop substrate color development. A 5% bovine serum albumin (BSA) block solution was prepared by dissolving 5.0 grams bovine serum albumin (98% purity, Fisher Scientific Inc.) in 100 mL PBS buffer.
The raw eggs (large size, grade A) used in this work were purchased from a local grocery store (United Supermarkets). The eggs were washed with DI water before being used as experimental materials.
Magnets
Neodymium disc magnets (N50 magnet, 1" diameter × 1/2" thick, nickel plated) from K&J Magnetics, Inc. (Plumsteadville, PA, USA) were used in this work to separate Salmonella-trapped MMP from egg sample matrix in the separation process. Each magnet was sealed inside a 1” × 2.5” rectangular plastic zip bag. A stainless steel wire was attached to the magnet in order to hang the magnet in the middle of egg sample mixture in a beaker. A DynaMagTM-5 magnet device from Life Technologies, Inc. (Carlsbad, CA, USA) was used to separate Salmonella-trapped MMP from aqueous solutions in the ELISA process. This magnetic device is suitable for separating MMP in small volume (<10 mL) solutions in test tubes.
Instruments
A mechanical blender purchased from a local grocery store was used to homogenize raw egg samples. A scanning UV/Vis spectrometer (Model UV2450, Shimadzu Corp. Kyoto, Japan) was used in this work to measure optical absorbance of colored solution obtained from ELISA process.
Procedure
Separating Salmonella bacteria from raw egg sample: A washed raw shell egg was broken and the materials within the shell were collected in a blender cup. The raw egg materials were homogenized by blending for 2 minutes. One hundred milliliters of DI water were used to wash the blender cup. The homogenized egg material and washing solution were transferred to a 400 mL glass beaker. DI water was further added to the homogenized raw egg material to make a 200 mL sample mixture. BacTrace® Anti-Salmonella CSA-1 MMP solution (0.10 mL) were added to the sample mixture. The obtained mixture was stirred for two minutes with a glass rod. The anti-Salmonella antibody coated MMP were then incubated with the sample mixture for 60 minutes. A N50 magnet (1” diameter) sealed in a plastic zip bag was placed into the center of sample mixture solution as showing in Figure 1, and stay in the solution for 60 minutes. The zip bag with the N50 magnet was then pulled out of the solution and transferred to a 50 mL glass beaker. The N50 magnet was removed from the zip bag and the plastic bag was zip-sealed. Five milliliters diluted PBS buffer solution were added to the beaker. The MMP were carefully removed from plastic bag surface by using a glass rod and re-dispersed into the PBS solution in the glass beaker by stirring the MMP/solution mixture with the glass rod. The MMP containing PBS solution was then transferred into a 10 mL glass test tube. The test tube was placed on the tube rack of the DynaMagTM 50 device to separate MMP from the solution. The PBS solution was discarded after 10 minutes the test tube was setting on the DynaMagTM 50 device. The Salmonellatrapped MMP in the test tube were washed three times with the diluted PBS buffer solution in order to remove egg sample matrix materials possibly absorbed onto the MMP. Each washing step consists of adding 5 mL diluted PBS buffer to the test tube, re-dispersing MMP to the PBS buffer solution by hand-shaking the MMP/PBS solution mixture in the test tube, incubating the MMP/PBS buffer solution for 5 minutes, separating MMP from PBS solution using DynaMagTM 50 device, and discarding the PBS buffer solution. The cleaned MMP were used for ELISA test.
Labeling trapped Salmonella bacteria with HRP-labeled anti- Salmonella antibody: Five milliliters 5% BSA solution were added to the test tube. The MMP were re-dispersed into the solution by shaking the test tube with the BSA solution. The BSA solution was incubated with the MMP in the test tube for 10 minutes, and then the MMP were separated from the solution by using the DynaMagTM 50 device. After discarding BSA solution, the MMP were washed three times with PBS buffer by following the washing procedure described above. Forty microliters of 1 μg/mL BacTrace® Anti-Salmonella, CSA-1 Antibody solution and 1.0 mL of PBS buffer solution were added to the test tube, and the MMP were re-dispersed to the mixture solution. The HRP-labeled anti- Salmonella antibody was incubated with the MMP for 20 minutes and the solution was discarded after separating the MMP with the DynaMagTM 50 device. The MMP and test tube were washed with PBS buffer three times.
Color development: SureBlue® TMB Microwell Peroxidase Substrate solution (0.60 mL) was added to the test tube and the MMP were re-dispersed by shaking the testing tube. The mixture was incubated for 10 min, and then 0.50 mL of the TMB BlueSTOPTM solution was added to stop the color development.
Measuring developed solution’s UV/Vis absorbance: The MMP were separated from the solution by using DynaMagTM 50 device. One milliliter of the colored solution was transferred from the test tube with a volumetric pipette to the 1-cm sample cuvette. One milliliter PBS buffer solution was added to the 1-cm cuvette in order to make the solution level in the cuvette high enough for optical absorbance measurement. The UV/Vis absorbance spectrum of the obtained solution in the wavelength range of 200 – 800 nm was recorded with the diluted PBS buffer as a reference solution, and the absorbance at 650 nm was used to calculate Salmonella typhimurium concentration in the raw egg sample.
Result and Discussion
Separating Salmonella typhimurium bacteria from raw egg sample matrix
The material inside the shell of a raw egg is a heterogeneous sample, contains yolk, egg white, chalaza, membranes. In order to make a homogeneous sample the contents inside the egg shell were first blended with a blender. The egg yolk and white contain high concentration of macro biomolecules, which makes the homogenized substance highly viscosity. It is difficult to evenly distribute MMP to a homogenized raw egg mixture. It is also difficult to pull up the MMP to the 1” diameter N50 magnet from an undiluted raw egg substance. Therefore, the egg materials were diluted with DI water to a 200 mL volume in order to reduce sample’s viscosity before adding antibody labeled MMP to the sample.
The BacTrace® Anti-Salmonella CSA-1 MMP was suggested to be used for separating Salmonella typhimurium bacteria from enriched food samples. The maker suggests adding 20 μL BacTrace® Anti-Salmonella CSA-1 MMP to 1 mL enriched sample solution for trapping the Salmonella bacteria. In this work the bacteria cells doped into one egg sample is in the same range (108 cells) as the number of the bacteria in 1 mL of an enriched sample. Therefore, if the incubation time is long enough, the number of antibody on MMP surface in 20 μL MMP solution should be enough to capture the bacteria cells from a sample mixture made from one egg in this work. In considering the fact that the sample volume is much large (200 mL) in this work, we choose to add 100 μL BacTrace® Anti-Salmonella CSA-1 MMP to a 200 mL egg sample mixture in order to reduce the MMP/sample incubation time.
Different approaches have been investigated for separating Salmonella-trapped MMP from raw egg sample mixture. These include: 1), placing the egg sample containing beaker on top of a 2” diameter N50 magnet disc in order to pull the Salmonellatrapped MMP to the beaker’ bottom; 2), directly placing a 1” diameter N50 magnet disc into a sample solution; 3), placing a 1” diameter N50 disc magnet into a plastic zip bag and deploying the sealed zip bag into the raw egg sample mixture in a beaker. Among these, it is difficult to separate egg materials from the MMP by using methods 1 and 2 because the egg sample matrix precipitated on the bottom of the beaker or absorbed onto the magnet’s surface together with MMP. The egg matrix materials cause interference in the ELISA process. Method 3 was adopted in this work because it can achieve the goal of separating Salmonella-trapped MMP from raw egg sample matrix. In this method, the plastic zip bag surface is hydrophobic, which avoided the absorption of raw egg sample matrix (hydrophilic macro biomolecules) onto its surface in separating Salmonella-trapped MMP from egg sample mixture.
It is possible that small quantity of egg sample matrix can be adsorbed onto the MMP surface. After separating Salmonellatrapped MMP from egg sample mixture, the MMP were washed with PBS buffer as described in the experimental section. The optical absorbance spectrum of the washing PBS buffer solution after separating MMP was measured and used as an indicator of washing efficiency. Figure 2 shows the UV/Vis absorption spectra of the washing PBS buffer solutions. The peak absorbance at around 270 nm indicates how much egg matrix material was washed off MMP into the PBS solution. It is obvious from the test results that most absorbed egg matrix substances were washed off the MMP after three washing steps.
Figure 2 Optical absorption spectra of PBS solutions obtained from washing Salmonella-trapped MMP, which have been incubated with a 0.05% triton X-100 diluted raw egg sample. The decrease of absorbance at around 270 nm with the number of washing indicates that egg matrix material was washed off MMP.
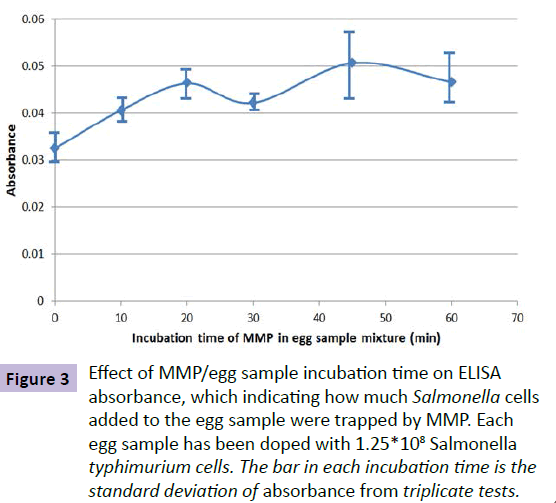
The time needed for anti-Salmonella typhimurium antibody on MMP to react with Salmonella typhimurium in an egg sample mixture was investigated. In this experiment, each egg sample solution was doped with 1.25*108 Salmonella typhimurium cells after homogenized and diluted with DI water. BacTrace® Anti-Salmonella CSA-1 MMP (0.10 mL) were then added to the egg samples. The time for incubating the MMP in each sample varied from 10 to 60 minutes. The Salmonella-trapped MMP were separated from sample matrix, cleaned and measured by using the ELISA process as described in the experimental part of this paper. The measured UV/Vis absorbance was used as a parameter for indicating how much Salmonella bacteria were captured by the anti-Salmonella antibody coated MMP. Figure 3 shows the relationship of obtained absorbance and MMP/egg sample incubation time. From this test result, 60 min was chosen as the MMP/egg sample mixture incubation time. It has to be mentioned that the incubation time can be significantly reduced if an appropriate device can be used to stir the MMP/sample mixture.
Figure 3 Effect of MMP/egg sample incubation time on ELISA absorbance, which indicating how much Salmonella cells added to the egg sample were trapped by MMP. Each egg sample has been doped with 1.25*108 Salmonella typhimurium cells. The bar in each incubation time is the standard deviation of absorbance from triplicate tests.
Figure 4 is the experimental results of investigating the time needed for the 1” diameter N50 magnet to pull up the MMP from 200 mL raw egg sample mixture. Each egg sample was doped with 1.25*108 Salmonella typhimurium cells. BacTrace® Anti- Salmonella CSA-1 MMP (0.10 mL) were incubated in the sample solution for 60 min. A N50 magnet (1” diameter) sealed inside a plastic zip bag was then deployed to the sample mixture as described in the experimental section of this paper. The magnet was positioned in the middle of the solution as possible. The incubation time of the magnet inside the sample solution varied for each sample. After pulled magnet out of the solution, the MMP collected from the solution on to the plastic zip bag surface was washed and analyzed with the described ELIZA method. The obtained absorbance value can be used as an indicator of how much MMP have been collected onto the plastic zip bag surface by the N50 magnet. This test result indicates that it take at least 60 minute for the 1” diameter N50 magnet to pull up all the MMP.
Figure 4 Effect of incubation time of the 1” diameter N50 magnet in egg sample mixture for pulling up Salmonella-trapped MMP, which is indicated by measured ELISA’s optical absorbance. Each egg sample has been doped with 1.25*108 Salmonella typhimurium cells. The bar in each incubation time is the standard deviation of absorbance from triplicate tests.
IMS-ELISA method
An ELISA method recommended by KPL Inc., the commercial company fabricating the ELISA reagents used in this work, was adopted and revised for detecting Salmonella typhimurium bacteria trapped onto the MMP. After trapping the Salmonella bacteria, the MMP were separated from raw egg matrix, transferred to a glass test tube and washed three times with the diluted PBS buffer solution. Five milliliters of the 5% BSA solution were added to the MMP in the test tube to block active sites on the surface of MMP particle as well as the glass test tube. The MMP and test tube were then washing three times with the diluted PBS buffer. The revised ELISA process includes labeling the Salmonella bacteria trapped on MMP surface with HRP-labeled anti-Salmonella antibody by incubating the MMP with 40 μL of 1.0 μg/mL HRP-labeled CSA-1 Antibody solution, three times PBS buffer washing. Each washing process consists of 5 minutes incubation with 5 mL PBS buffer solution, separating MMP from PBS buffer solution with the DynaMagTM-5 magnet device, and discarding PBS buffer solution.
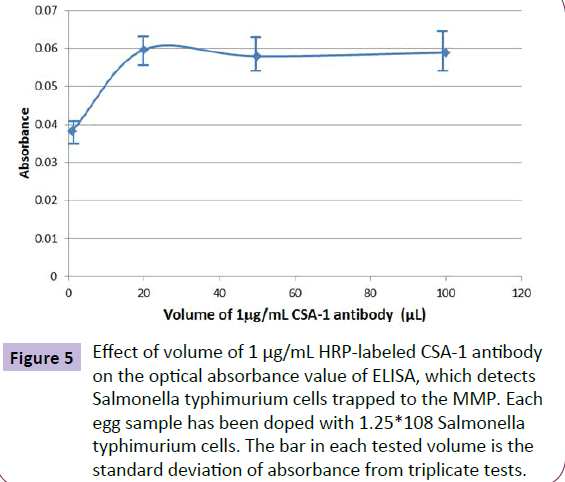
The effect of HRP-labeled antibody (CSA-1) quantity on ELISA absorbance signal was investigated by adding different volume of the 1.0 μg/mL antibody solution to label Salmonella typhimurium bacteria trapped on MMP. Figure 5 shows the experimental results. The optimized volume of the 1.0 μg/mL HRP-labeled antibody solution is 40 μL.
Figure 5 Effect of volume of 1 μg/mL HRP-labeled CSA-1 antibody on the optical absorbance value of ELISA, which detects Salmonella typhimurium cells trapped to the MMP. Each egg sample has been doped with 1.25*108 Salmonella typhimurium cells. The bar in each tested volume is the standard deviation of absorbance from triplicate tests.
The MMP in the test tube was then incubated with 0.60 mL SureBlue substrate solution for 10 minutes. The SureBlue stop solution (0.50 mL) was then added to stop color development. MMP were then separated from the colored solution with the DynaMagTM-5 magnet device. The colored solution was transferred to a 1.0 cm quartz cell for UV/Vis absorption spectrometric analysis. The solution’s absorption spectrum was recorded. The absorbance at 650 nm was used for detecting the number of Salmonella cells in the raw egg sample.
Figures of merit of the developed method
In order to establish a calibration curve for the developed method, different number of Salmonella typhimurium cells were added to raw egg samples, and the obtained samples were analyzed with the developed method. The obtained relationship of Salmonella typhimurium cell number in one egg sample with the measured absorbance value is illustrated in Figure 6. Six raw egg samples without Salmonella typhimurium doping were analyzed with the developed procedure. The standard deviation of absorbance value of these blank samples was calculated to be 0.0007. The detection limit of the developed method, which is defined as the number of Salmonella typhimurium cells in one egg sample gives an absorbance value equals three times of the standard deviation of blank sample’s absorbance value, is calculated to be 1.4*107 Salmonella typhimurium cells/egg.
Conclusions and Further Work
The feasibility of using IMS for directly separating Salmonella typhimurium cells from raw egg samples was investigated. Experimental results indicate that this simple method can be used to separate Salmonella bacteria from complex raw egg samples. When an ELISA method with UV/Vis absorption spectrometry was employed for detecting the separated bacteria, the developed method can detect 1.4*107 Salmonella typhimurium cells in one raw egg within 5 hours. More sensitive detection methods, such as laser induced fluorescence [3-5], chemiluminescence [20], and liquid core waveguide fiber optic spectrometry [21,22] will be investigated for detecting separated Salmonella bacteria. With the adaptation of more sensitive detection technologies, it is expected that a quick method can be developed for directly detecting single digit number of Salmonella cells in a raw egg sample [23,24].
Acknowledgement
This work was supported by National Institute of Food and Agriculture, United States Department of Agriculture through award number: 2012-69003-19620.
References
- Dwivedi HP,Jaykus LA (2011) Detection of pathogens in foods: the current state-of-the-art and future directions.Critical Rev Microbiol 37: 40-63.
- Nugen SR, Baeumner AJ (2008)Trends and opportunities in food pathogen detection. Anal Bioanal Chem391: 451-454.
- Moerner WE, Fromn DP (2003) Methods of single molecular fluorescence spectroscopy and microscopy. Rev SciInstrum 74: 3597-3619.
- Ishijima A, Yanagida T (2001) Single molecule nanobioscience. TRENDS in Biochem Sci 26: 438-444.
- Benito A, Garcia-Canas V, Alejandro C, Ramon G, Rosa A (2004) Simultaneous and sensitive detection of three foodborne pathogens by multiplex PCR, capillary gel electrophoresis, and laser induced fluorescence. J Agr Food Chem 52: 7180-7186.
- Giebel R, Worden C, Rust SM, Kleinheinz GT, Robbins M, et al. (2010) Microbial fingerprinting using matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS): applications and challenges.AdvAppl Microbiol 71: 149-184.
- USDA Food Safety and Inspection Service, Microbiology Laboratory Guidebook.
- US Food and Drug Administration. Bacteriological Analytical Manual (BAM).
- Kendall P, Payton L (2008) Cost of preserving and storing food. Food and Nutri Ser No 8: 704.
- Stevens KA, Jaykus LA (2004) Bacterial separation and concentration from complex sample matrices: a review.Critical Rev Microbiol 30: 7-24.
- Benoit PW, Donahue DW (2003) Methods for rapid separation and concentration of bacteria in food that bypass time-consuming cultural enrichment.J Food Protect 66: 1935-1948.
- SafarikI, Safarikova M (1999) Use of magnetic techniques for the isolation of cells. J Chromatogr B 722: 33-53.
- Mauro M, Luca G, Bruce IJ (2006)The use of magnetic nanoparticles in the development of new molecular detection systems. J NanosciNanotechnol 6: 2302-2311.
- Steingroewer J, Knaus H, Bley H, Boschke E (2005) A rapid method for the pre-enrichment and detection of Salmonella typhimurium by immunomagnetic separation and subsequent fluorescence microscopical techniques.Eng Life Sci 5: 267-272.
- Liu Y, CheY, Li Y (2001) Rapid detection of Salmonella typhimurium using immunomagnetic separation and immune-optical sensing method.SensActuat B72: 214-218.
- Guo PL, Tang M, Hong SL, Yu X, Pang DW, et al. (2015) Combination of dynamic magnetophoretic separation and stationary magnetic trap for highly sensitive and selective detection of Salmonella typhimurium in complex matrix. Biosens Bioelectron 74: 628-636.
- Chen WY, Chen YC (2006) Affinity-based mass spectrometry using magnetic iron oxide particles as the matrix and concentrating probes for SALDI MS analysis of peptides and proteins. Anal BioanalChem 386: 699-704.
- Aguilar-Arteaga K, Rodriguez JA, Barrado E (2010) Magnetic solids in analytical chemistry: A review. Anal ChimActa 674: 157-165.
- Cudjoe KS, Hagtvedt T, Dainty R (1995) Immunomagnetic separation of Salmonella from foods and their detection using immunomagnetic particle (IMP) –ELISA. Intern J Food Microbiol 27: 11-25.
- RijpensN, HermanL, VereeckenF, JannesG, De SmedtJ, et al.(1999) Rapid detection of stressed Salmonella spp. in dairy and egg products using immunomagnetic separation and PCR. Intern J Food Microbiol 46: 37-44.
- Siriken B, Turk H, Yildirim T, Durupinar B, ErolI (2015) Prevalence and characterization of Salmonella isolated from chicken meat in Turkey. J Food Sci 80: 1044-1050.
- RodaA, Pasini P, Mirasoli M, Michelini E, Guardigli M (2004) Biotechnological applications of bioluminescence and chemiluminescence. TRENDS in Biotechnol 22: 295-303.
- Le T,Tao S (2011) Intrinsic UV absorption spectrometry observed with a liquid core waveguide as a sensor technique for monitoring of ozone in water. Analyst 136: 3335-3342.
- Tao S, Winstead CB, Xian H, Krunal S (2002) A highly sensitive hexachromium monitor using water core optical fiber with UV LED. J Environ Monitor 4: 815-818.
Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences